- All Products
- MonoliF - Monolignol Fluorescent kit
MonoliF - Monolignol Fluorescent kit
https://www.idylle-labs.com/shop/monolif-monolignol-fluorescent-kit-840 https://www.idylle-labs.com/web/image/product.template/840/image_1920?unique=d0d5e10Kit content : 1) Monolignol reporters (S-MonoliF and/or G-MonoliF), corresponding to 300mL of working solution. 2) Solution for the detection of monolignol with click chemistry, available in small or large kits.
A technology developed by Clémence Simon, Christophe Biot and Cédric Lion (Unit of Structural and Fonctional Glycobiology (UGSF), UMR 8576, France)
MonoliF is a labeling kit designed to study lignin biosynthesis inside plants. This kit uses monolignol reporters that are incorporated into lignin just like naturally occuring monolignols. G and S MonoliF can then be detected via fluorescence after labeling with click chemistry. It is the only method that will allow you to study lignin composition and localisation in a dynamic manner, since only the MonoliF (and not naturally occuring monolignols) will be labeled.
Main features of MonoliF

Biological relevance
Modifications on S and G MonoliF cannot be detected by living cell, meaning they are seen as naturally occuring monolignols by plants.

Deeper insights on lignin
You can analyze which type of monolignol is integrated, its localisation, as well as its dynamics.

Live imaging
Lignin dynamics can be studied in seedlings, live stem or cross section of plants (using confocal microscope or airyscan)
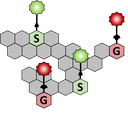
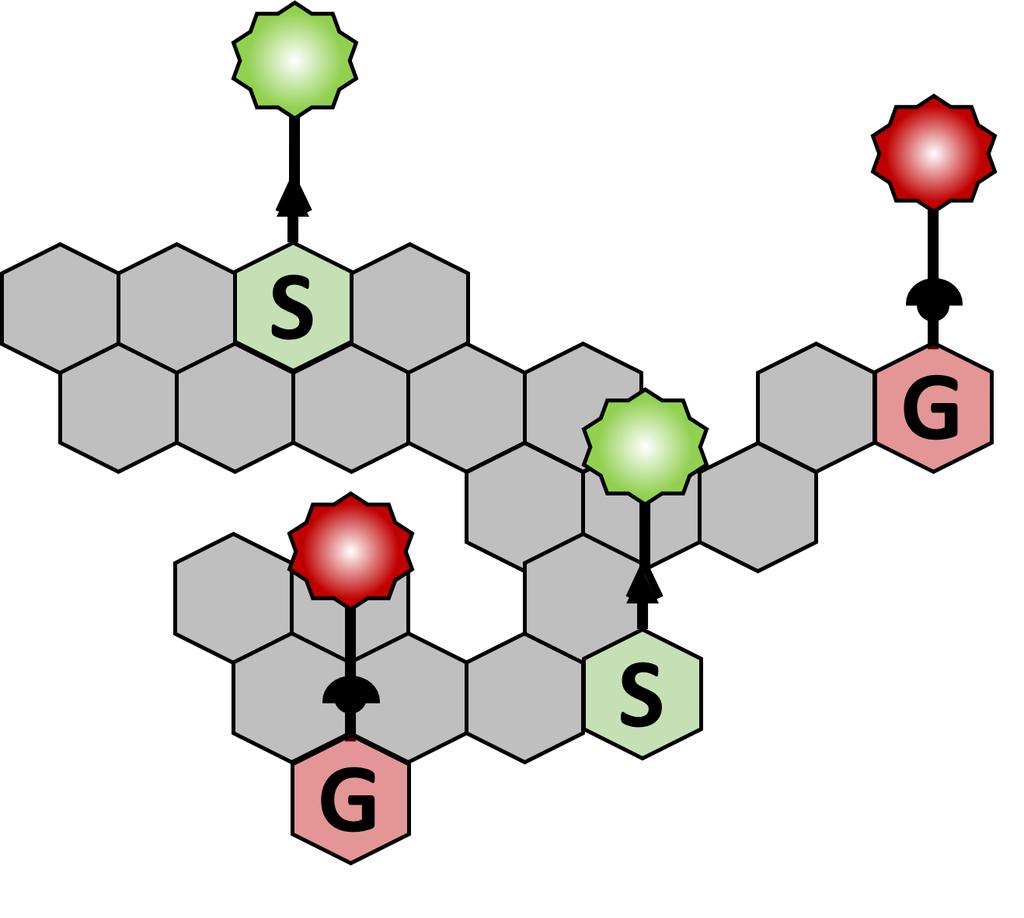
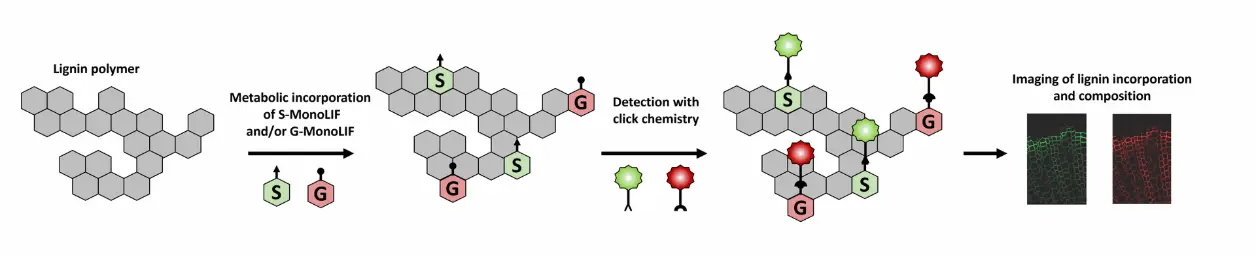
How does MonoliF kit work ?
- S and/or G MonoliF analogs bearing either an azide or an alkyne group are metabolically incorporated into lignifying cell walls.
- Detection of MonoliF is performed with click chemistry kit in a straightforward, bioorthogonal manner (the reaction does not interact with natural biochemical processes).
- Lignin incorporation and composition can be studied with fluorescence signal.
Kit content of MonoliF

*One experiment corresponds to 300µL.
Applications
Type of sample : MonoliF has been already validated on cross sections, stems, and seedlings.
Type of plants MonoliF has been tested on Arabidospis Thaliana, Poplar, Tobacco and flax.
Specifications of MonoliF
Fluorescence and microscopy :
- Tagged G-MonoliF : Abs/Em = 546/565 nm
- Tagged S-MonoliF : Abs/Em = 501/526 nm
We recommand using a confocal microscope or an airyscan
Storage and stability :
- Store G and S MonoliF at -20°C. Stable up to 4 years
- Store Click chemistry fluorophores at -20°C. Stable up to 1 year.
Additional ressources
Follow of cell-specific monolignol reporter incorporation with MonoliF* (1)
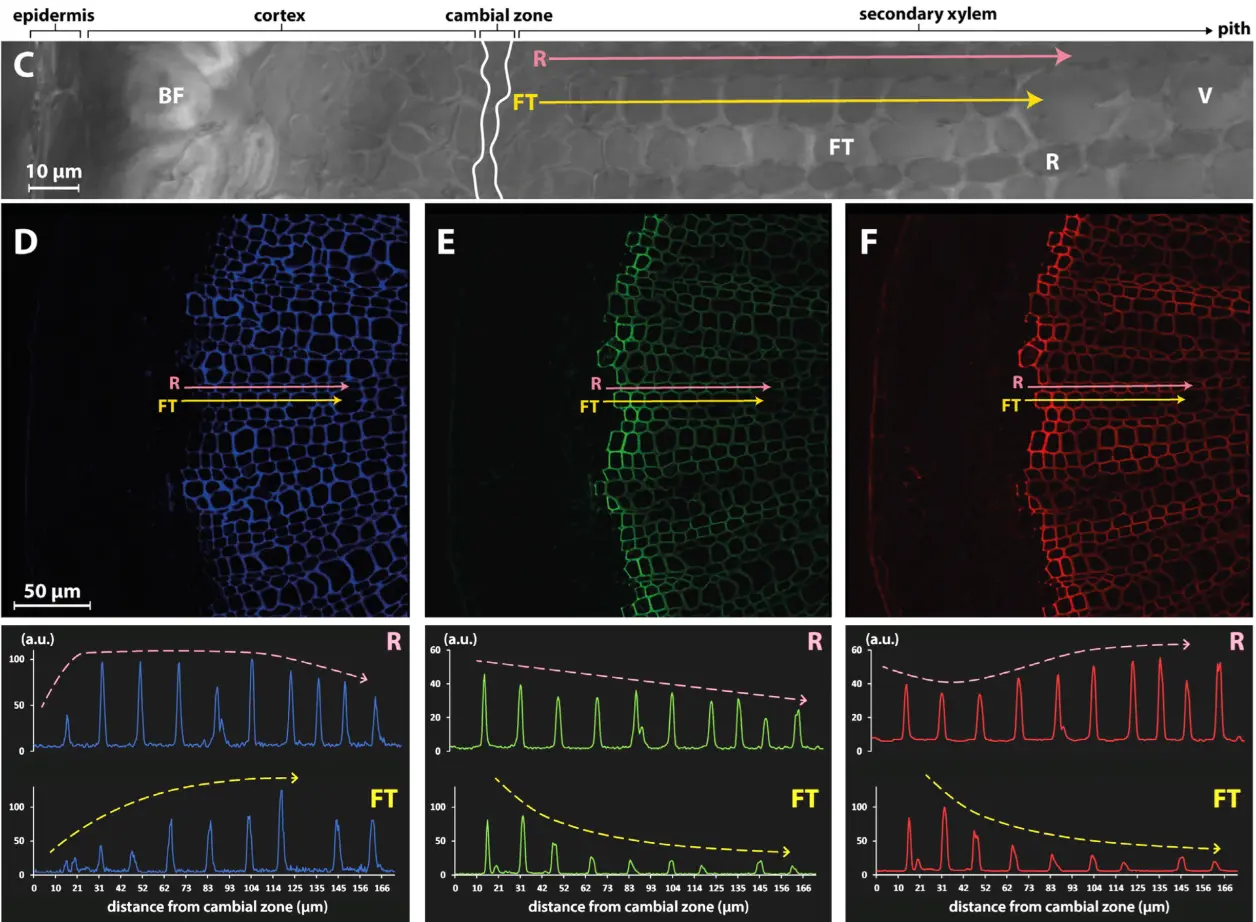
Observation of monolignol incorporation (H monolignol in green, G monolignol in red) in a cross section of flax stem. The incorporation of monolignol varies across the distance from the cambial zone but also according to the type of cell (Ray cells, R, pink arrow; Fiber tracheid cell, FT, yellow arrow)
Versatility of MonoliF labeling across multiple plant types (2)
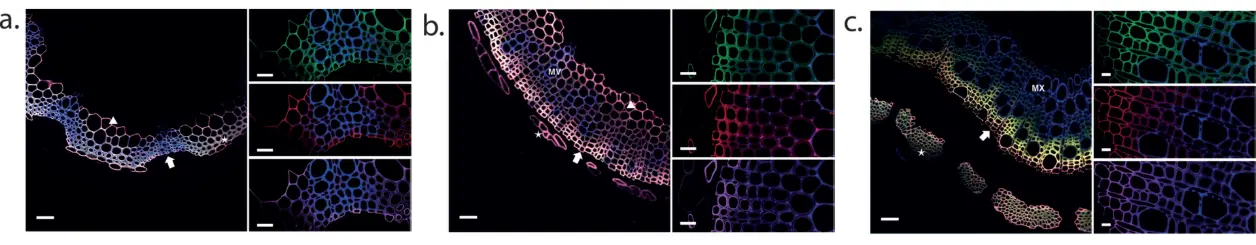
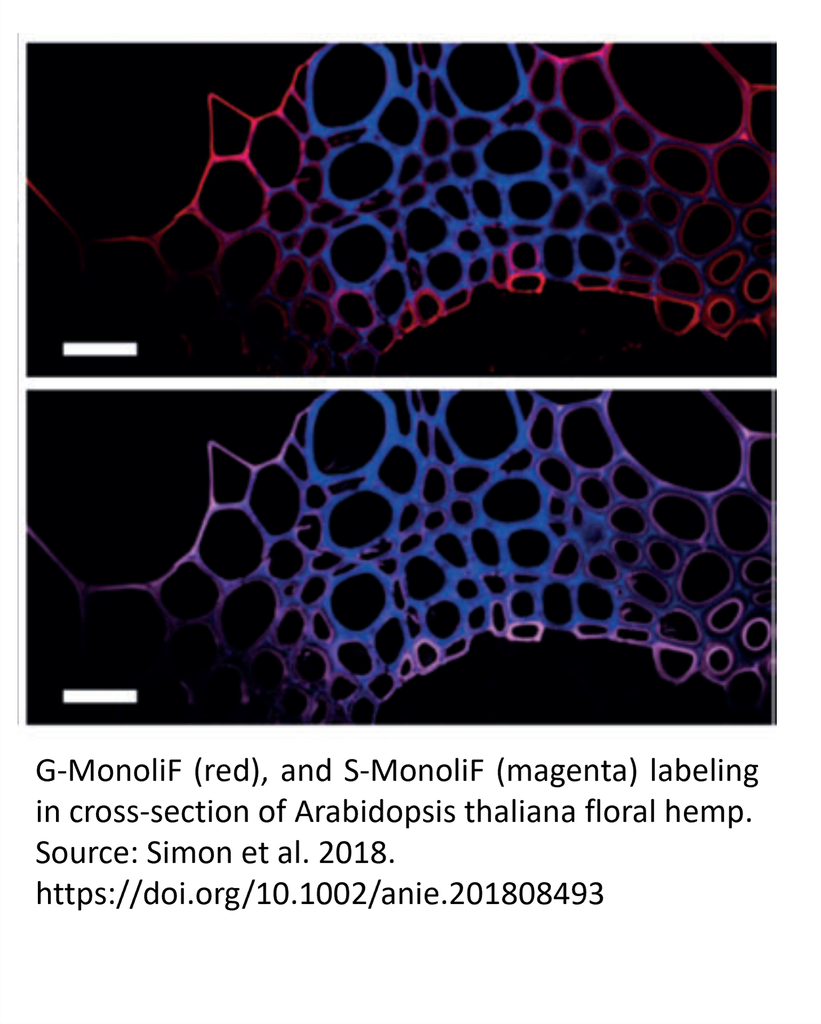
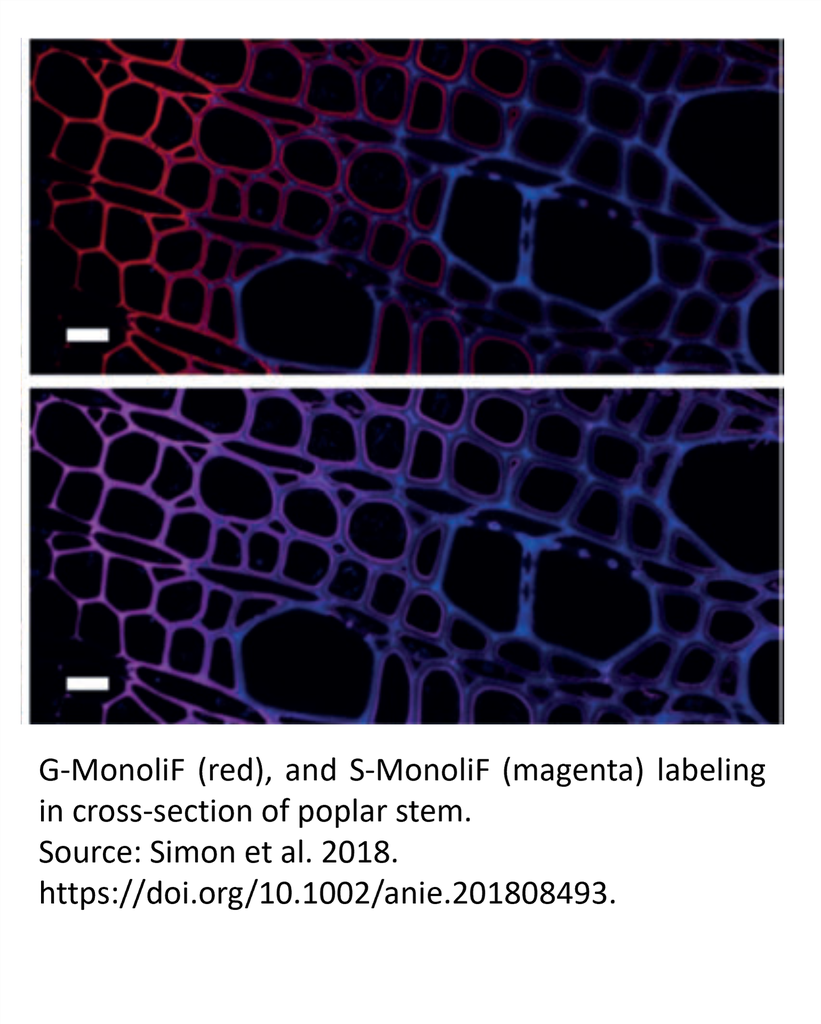
Incorporation of H (green), G (red), and S (magenta) monolignols in cross-section of Arabidopsis thaliana floral hemp (a), tobacco stem (b) and poplar stem (c).
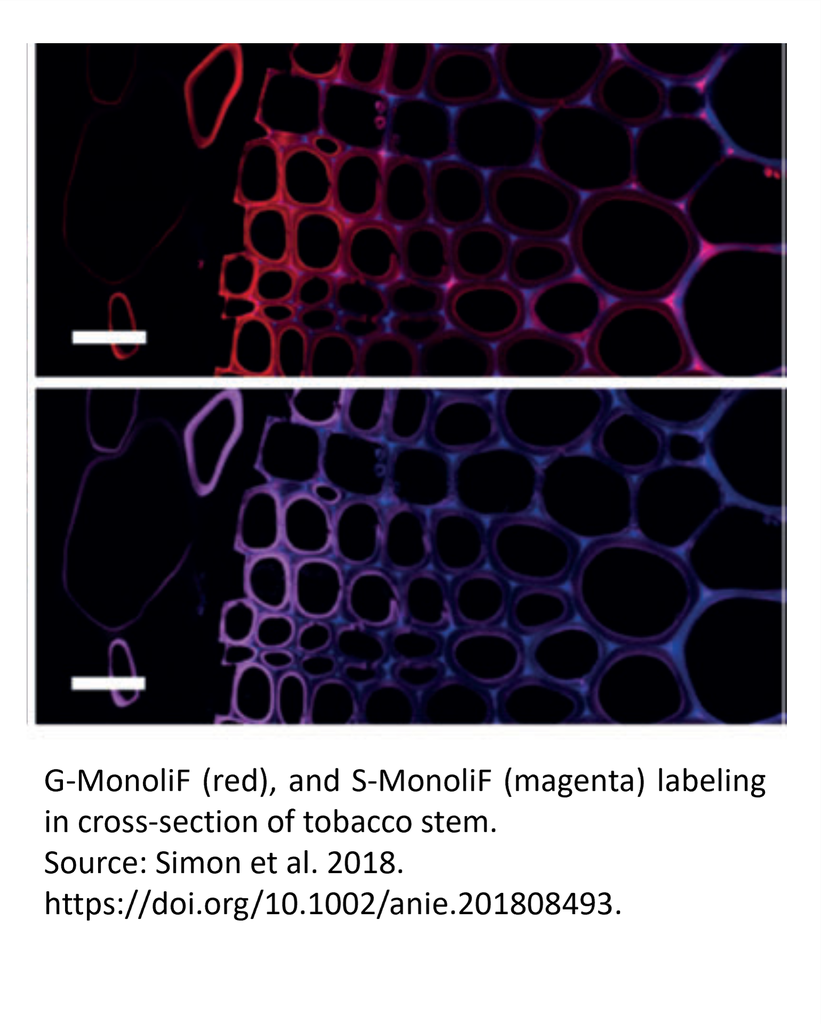
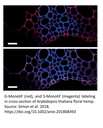
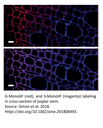
Labeling of G and S monolignols in flax stem cross section with MonoliF* (2)

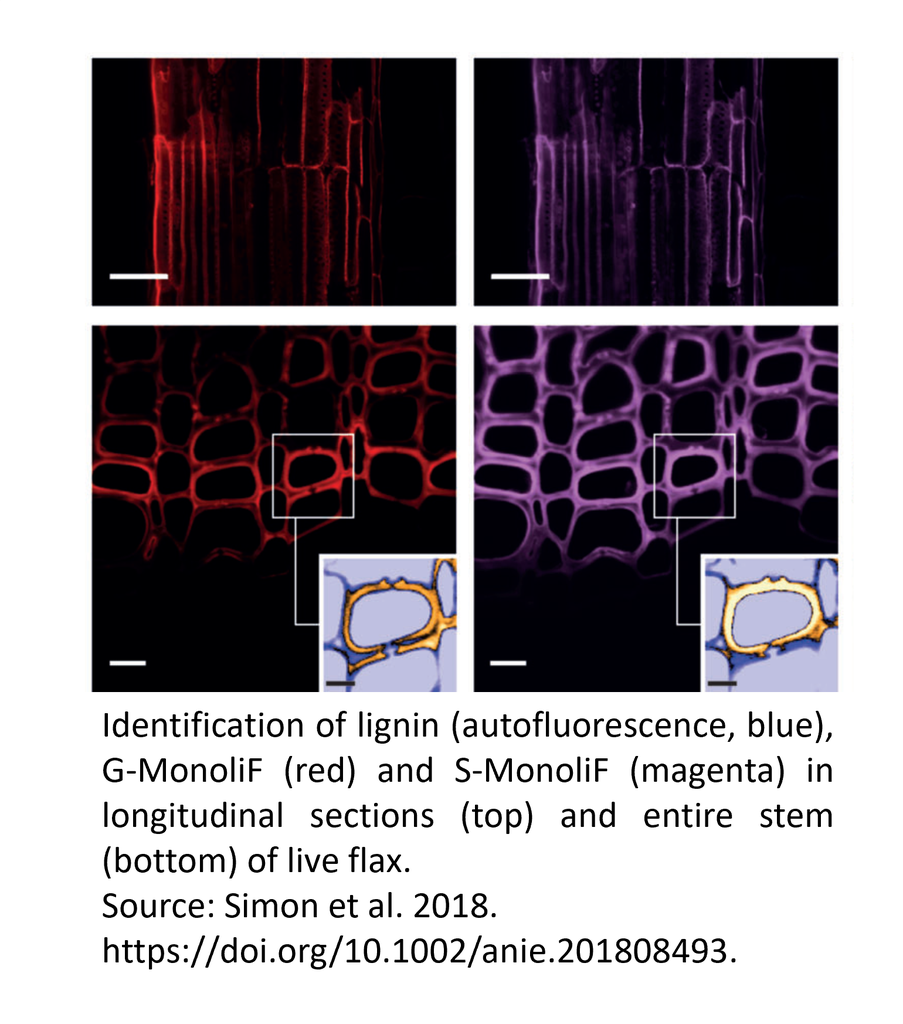
Identification of lignin (autofluorescence, blue), G monolignol (red) and S monolignol (magenta) in transversal cross-sections (a), longitudinal sections (b) and entire stem (c) of live flax. Scale bar=50 mm (a,b), 10 mm (c) and 5 mm (insert).
Analysis of flax xylem pit border region with MonoliF*

Airyscan images of the 3 monolignols reporters : H (in blue), G (in red) and S (in red) BLISS monolignols (A). The merged image shows cell walls and a bordered pit (BP) between two cells in flax xylem (B). Orange lines P1−P3 show vector lines used to trace individual reporter (H, G, S) fluorescence intensity profiles along the lines plotted in (C). Scale bar = 3 μm.
References :
1. Lion et al., Cell Chemical Biology, 2017 http://dx.doi.org/10.1016/j.chembiol.2017.02.009
2. Simon et al., Angew. Chem. Int. Ed. 2018. https://doi.org/10.1002/anie.201808493.
3. Simon & al 2023, Chem Biomed Imaging, https://doi.org/10.1021/cbmi.3c00052
* Experiments carried out by researchers were performed using varying monolignol analogs. Please refer to the publications for detailed informations.
Check out the Publications section for additional results.
For dual click chemistry labelling, two types of click reactions are employed: SPAAC (Strain-Promoted Azide–Alkyne Cycloaddition), which proceeds without a catalyst will be performed first. It will be followed by CuAAC reaction (Copper-Catalyzed Azide–Alkyne Cycloaddition), which requires copper as a catalyst. These reactions are highly specific and efficient, forming stable triazole linkages under mild conditions. The click chemistry labelling is bioorthogonal, and reliable, making it ideal for labeling and bioconjugation in complex biological systems.
The signal is very specific. Click chemistry is designed so that azide and alkyne groups only react with each other and do not interact with any other biological molecules. This ensures that the labelling marks only the target molecules. Importantly, azide and alkyne do not interact to each other unless specific conditions (catalyser or special form of alkyne) are present. Therefore, S and G MonoliF will not bind to each other.
No. Only the MonoliF, which contain an alkyne or an azide group will be labeled.
These molecules are detected by click chemistry, which makes them fluorescent after labeling.
You can compare fluorescence intensity between samples to get relative quantification. This means you can measure differences between conditions, but not necessarily get absolute molecular amounts.
It is recommended to use confocal microscopy. Confocal microscopes provide better resolution than standard inverted epifluorescence microscopes. You may also use an Airyscan detector, which improves the signal-to-noise ratio and the spatial resolution. With a non-confocal microscope, accurate quantification of the signal is generally not possible.
No, it is not toxic. The azide and alkyne groups on the monolignol analogs do not interfere with biological processes, and the click chemistry reaction itself is biorthogonal, meaning it works independently of normal cellular chemistry. In one publication (https://doi.org/10.1002/anie.201808493, Supplementary data), flax seeds were incubated with an analog similar to S-MonoliF for 7 days, and no difference in plant architecture was observed.
The team that originally developped MonoliF have published several papers using their innovation (previously reported as BLISS) :
BLISS: A Bioorthogonal Dual-Labeling Strategy to Unravel Lignification Dynamics in Plants Cell Chemical Biology, 2017. Cédric Lion, Clémence Simon, Brigitte Huss, Anne-Sophie Blervacq, Louis Tirot, Djadidi Toybou, Corentin Spriet, Christian Slomianny, Yann Guerardel, Simon Hawkins, Christophe Biot. (https://doi.org/10.1016/j.chembiol.2017.02.009)
BLISS: Shining a light on lignification in plants, Plant Signaling and Behavior, 2017. Clémence Simon, Cédric Lion, Brigitte Huss, Anne-Sophie Blervacq, Corentin Spriet, Yann Guérardel, Christophe Biot and Simon Hawkins. (https://doi.org/10.1080/15592324.2017.1359366 )
One, Two, Three: A Bioorthogonal Triple Labelling Strategy for Studying the Dynamics of Plant Cell Wall Formation In Vivo, Angewandte Chemie International Edition, 2018. Clémence Simon, Cédric Lion, Corentin Spriet, Fabien Baldacci-Cresp, Simon Hawkins, Christophe Biot.(https://doi.org/10.1002/anie.201808493)
EPR imaging of sinapyl alcohol and its application to the study of plant cell wall lignification, Chemical Communications, 2020. Clémence Simon, Cédric Lion, Hania Ahouari, Hervé Vezin, Simon Hawkins, Christophe Biot. (https://doi.org/10.1039/D0CC05218C)
REPRISAL: mapping lignification dynamics using chemistry, data segmentation, and ratiometric analysis, Plant Physiology, 2022. Oriane Morel, Cedric Lion, Godfrey Neutelings, Jonathan Stefanov, Fabien Baldacci‐Cresp, Clemence Simon, Christophe Biot, Simon Hawkins, Corentin Spriet. (https://doi.org/10.1093/plphys/kiab490)
Exploring Lignification Complexity in Plant Cell Walls with Airyscan Super-resolution Microscopy and Bioorthogonal Chemistry, Chemical and Biomedical Imaging, 2023. Clémence Simon, Oriane Morel, Godfrey Neutelings, Fabien Baldacci-Cresp, Marie Baucher, Corentin Spriet, Christophe Biot, Simon Hawkins, Cédric Lion. (https://doi.org/10.1021/cbmi.3c00052)