- All Products
- PhaseCapture

PhaseCapture
https://www.idylle-labs.com/shop/phasecapture-970 https://www.idylle-labs.com/web/image/product.template/970/image_1920?unique=b5e7067DNA/RNA extraction kit for phase-separated nuclear compartments
A technology developed by Thierry Forne and Cosette Rebouissou (Institut de Génétique Moléculaire de Montpellier - CNRS UMR 5535, France)
The PhaseCapture kit isolates membrane-less compartments present in the cell nuclei while perfectly preserving their nucleic acid contents. It is used to recover and analyze the DNA/RNA associated with all nuclear bodies and large ribonucleoprotein (RNP) complexes formed by phase separation in the cell nucleus.
High specificity
Nuclear bodies get specifically isolated thanks to high-salt treatments before extracting their associated DNA/RNA contents. No risk of biases due to random, aspecific interactions as it can be the case with chemical crosslinking procedures.
Wide coverage
Relies on phase properties, rather than specific protein targets, to get an exhaustive sampling of all genomic sequences associated with nuclei sub-compartments.
Simple protocol
No need to go through the generation of transgenic cell lines or other fastidious protocols. PhaseCapture offers the first ready-to-use kit for efficient purification of nuclear bodies for genetic analyses.
Applications
Extracted contents: DNA & RNA
Kit specificity: all membrane-less compartments formed by liquid-liquid phase separation in the cell nucleus, including histone locus bodies (HLBs), PML bodies, Cajal bodies, nuclear speckles & paraspeckles, nucleoles* & transcriptional condensates.
*including perinucleolar heterochromatin
Starting material: isolated nuclei from human & murine cells/tissues
Validated samples:
- mouse ESCs & ESC-derived neurons
- human fibroblasts (IMR-90), lymphocytes, neurons and iPS-derived motoneurons
Compatible assays:
- DNA isolation for PCR, qPCR or sequencing*
*validated sequencing method: Illumina HiSeq 2000 (NGS sequencing)
- RNA isolation for RT-qPCR or RNA-seq
How does it work?
The HRS-Seq method relies on a specific high-salt treatment of cell nuclei that makes all nuclear bodies formed by liquid-liquid phase separation insoluble while perfectly preserving their nucleic acid contents.
All DNA/RNA associated with nuclear bodies from the same sample (insoluble fraction) are then separated from the rest of the genome (soluble fractions) using simple enzymatic digestion and washing steps. They can be further analyzed using PCR, qPCR, RT-qPCR or sequencing.
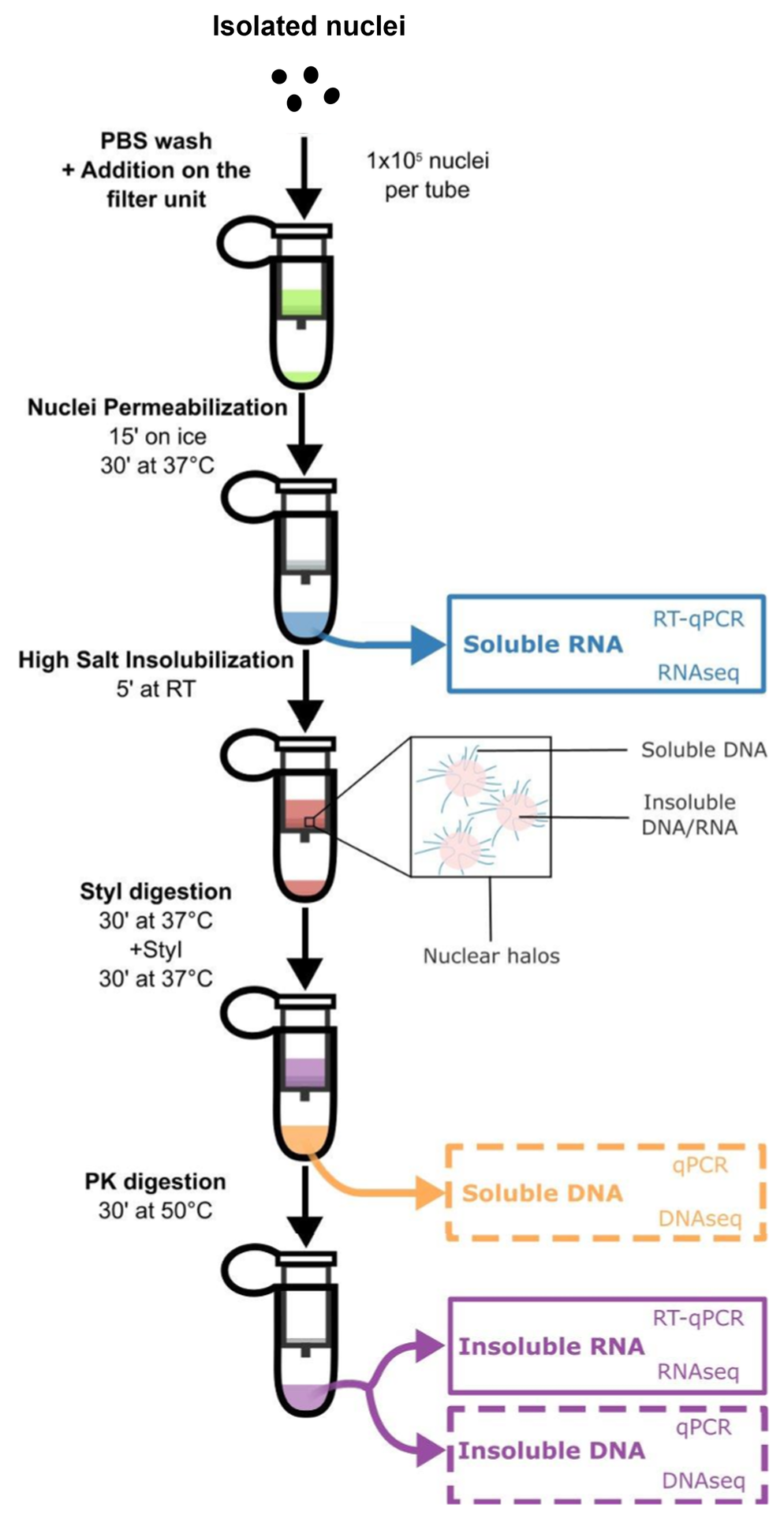
Figure modified from Lecouvreur N. et al, Methods in Molecular Biology, 2025, 2962, pp.141-151.
Use cases
- Exploring the role of nuclear bodies & phase separation in epigenetics (Baudement et al, 2018)
- Assessing nuclear body changes in response to external stresses at the genomic & transcriptomic level (i.e. heterochromatin changes, RNA composition, splicing profile)
- Identifying gene contents of nuclear bodies & obtain a global profiling of associated sequences
Filtration kit
The filtration kit contains everything you need to process your nuclei samples (6 samples per run are extracted in parallel ). It is reusable and only needs to be purchased once for infinite extraction assays.
Kit contents:
Steel air valves x6
O-ring x6
Plugs x6
Pump x1
Tubing
Small kit
The small extraction kit contains the buffers and filtration units required to extract DNA/RNA from isolated nuclei (up to 24 samples*).
- Kit contents:
Permeabilization buffer
Extraction buffer
Enzyme buffer
10X PK buffer
Filtration units x24
Large kit
The large extraction kit contains the buffers and filtration units required to extract DNA/RNA from isolated nuclei (up to 90 samples*).
- Kit contents:
Permeabilization buffer
Extraction buffer
Enzyme buffer
10X PK buffer
Filtration units x90
*For sequencing applications, we recommend pooling the DNA/RNA contents extracted from 6 separate filtration units to get enough material.
Chromosomal mapping of genomic regions associated with nuclear bodies
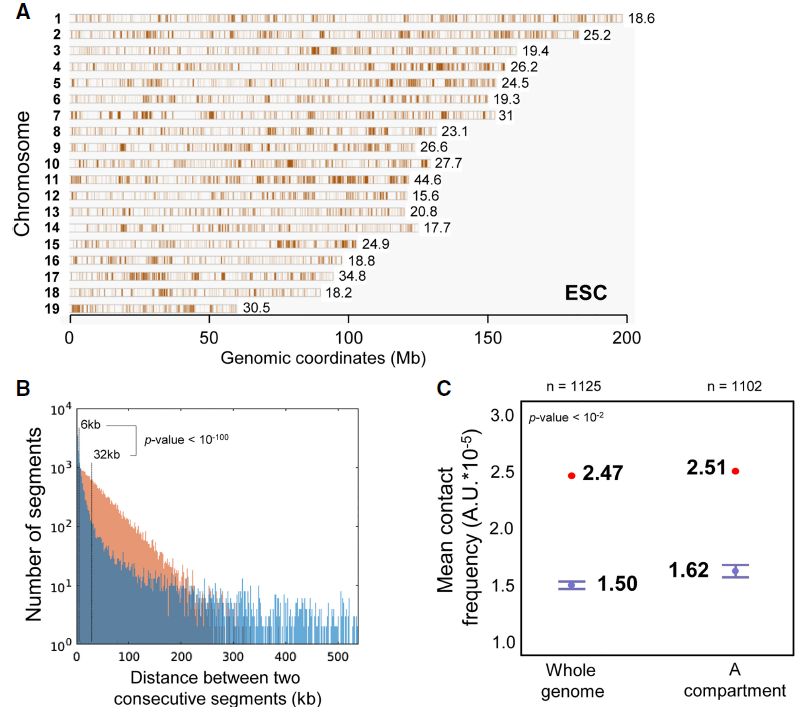
High-salt recovered sequences are associated with active epigenetic marks
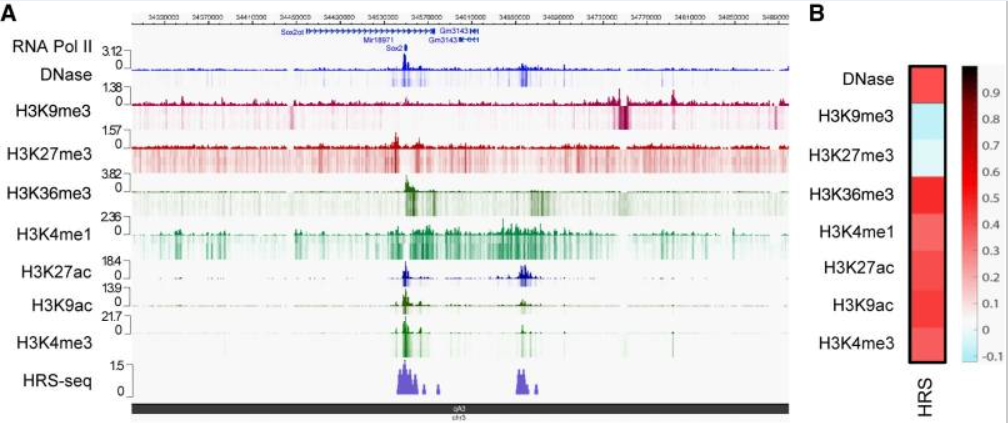
les design...
et leurs applications
Protocol coming soon
PhaseCapture is a general method that allows to insolubilize and purify all membrane-less compartments formed by liquid-liquid phase separation in the cell nucleus. They include Histone locus bodies (HLBs), PML bodies, Cajal bodies, nucleoles (incl. prinucleolar heterochromatin), nuclear speckles, paraspeckles and transcriptional condensates.
Yes, the extracted DNA and RNA contents can be further analyzed by sequencing (DNAseq or RNA-seq). Check the publication below for a concrete example of next-generation sequencing of DNA extracted using PhaseCapture (Illumina Hi-Seq 2000) and the detailed associated protocol from the library preparation to bioinformatic analyses:
Cosette Rebouissou, Marie-Odile Baudement, ThierryForné. The High-salt Recovered Sequence-sequencing (HRS-seq) method: exploring genome association with nuclear bodies. Methods in Molecular Biology, 2022, Spatial Genome Organization: Methods and Protocols, 2532,pp.187-197. 10.1007/978-1-0716-2497-5_9
The PhaseCapture kit is used to recover DNA/RNA contents from all membrane-less compartments formed by phase separation in the cell nucleus (incl. Histone locus bodies, PML bodies, Cajal bodies, nuclear speckles & paraspeckles, nucleoles & transcriptional condensates).
To recover DNA/RNA contents from specific compartments, the nuclear bodies of interest can be specifically inactivated using CRISPR/Cas9 or RNA interference to inhibit a factor that is essential to their formation. By extracting DNA/RNA contents using PhaseCapture from both WT and mutated samples in parallel, it is then possible to identify, by comparison, the regions associated with the body of interest.
We don’t recommend using PhaseCapture for protein isolation as the detergents present in the buffers might alter protein contents.
The various nuclear bodies and RNP complexes recovered using PhaseCapture were identified using a combination of qPCR and high-throughput sequencing to identify enriched genes associated with specific nuclear bodies, and immunofluorescence assays for direct imaging of specific markers in the insoluble fractions. Check the publication below for more detailed information.
Baudement MO, Cournac A, Court F, Seveno M, Parrinello H, Reynes C, Sabatier R, Bouschet T, Yi Z, Sallis S, Tancelin M, Rebouissou C, Cathala G, Lesne A, Mozziconacci J, Journot L, Forné T. High-salt-recovered sequences are associated with the active chromosomal compartment and with large ribonucleoprotein complexes including nuclear bodies. Genome Res. 2018 Nov;28(11):1733-1746. doi: 10.1101/gr.237073.118. Epub 2018 Oct 4. PMID: 30287550; PMCID: PMC6211644.
Yes, PhaseCapture can be used to assess nuclear body changes in response to external stresses at the genomic & transcriptomic level (i.e. heterochromatin changes, RNA composition, splicing profile). It was successfully used to extract DNA contents from senescent cells.
Lecouvreur N, Rebouissou C, Lesne A, Forné T. Simultaneous Profiling of Transcripts and Genomic Regions Associated with Nuclear Bodies Using the RNA-DNA High-Salt Recovered Sequence (RD-HRS) Method. Methods Mol Biol. 2025;2962:141-151. doi: 10.1007/978-1-0716-4726-4_10. PMID: 40699427.
Baudement MO, Cournac A, Court F, Seveno M, Parrinello H, Reynes C, Sabatier R, Bouschet T, Yi Z, Sallis S, Tancelin M, Rebouissou C, Cathala G, Lesne A, Mozziconacci J, Journot L, Forné T. High-salt-recovered sequences are associated with the active chromosomal compartment and with large ribonucleoprotein complexes including nuclear bodies. Genome Res. 2018 Nov;28(11):1733-1746. doi: 10.1101/gr.237073.118. Epub 2018 Oct 4. PMID: 30287550; PMCID: PMC6211644.
Lesne A, Baudement MO, Rebouissou C, Forné T. Exploring Mammalian Genome within Phase-Separated Nuclear Bodies: Experimental Methods and Implications for Gene Expression. Genes (Basel). 2019 Dec 17;10(12):1049. doi: 10.3390/genes10121049. PMID: 31861077; PMCID: PMC6947181.
Rebouissou C, Baudement MO, Forné T. The High-Salt Recovered Sequence-Sequencing (HRS-seq) Method: Exploring Genome Association with Nuclear Bodies. Methods Mol Biol. 2022;2532:187-197. doi: 10.1007/978-1-0716-2497-5_9. PMID: 35867250.